Latest ClickGene publications
As the ClickGene project is coming to an end, ClickGene early-stage-researchers (ESRs) continue to disseminate their work through publications in renowned Chemistry journals.
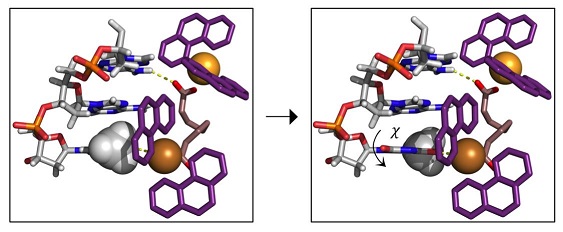
ESRs Nicolo Fantoni, Georgia Menounou and Gianluca Toniolo recently published a paper in Chemistry: A European Journal, which reports a new type of artificial metallo-nuclease for the building of robust inorganic scaffolds.
In November 2018, the work of Gianluca Toniolo (in collaboration with ESRs Maria Louka, Georgia Menounou and Nicolo Fantoni) has been published on the biological reactivity of a novel artificial nuclease. An article of interest for chemotherapeutic activities and published on the American Chemical Society journal ACS Omega.
The Kellett Group in DCU has also recently published two papers in Chem. Soc. Rev and ChemPlusChem. In “Molecular methods for assessment of non-covalent metallodrug–DNA interactions“, the Kellett Group describes a range of molecular methods to explore novel metallodrug-DNA interactions. In “Efficient DNA Condensation by a C3‐Symmetric Codeine Scaffold”, the authors propose a novel nucleic acid condensation agent to be used in gene therapy.
More ClickGene-related publications are foreseen within the next months. You can find the current list of journal articles on the publications page.